Genetic codebreaking on wheat years ahead of schedule
By Staff
| 3 min read
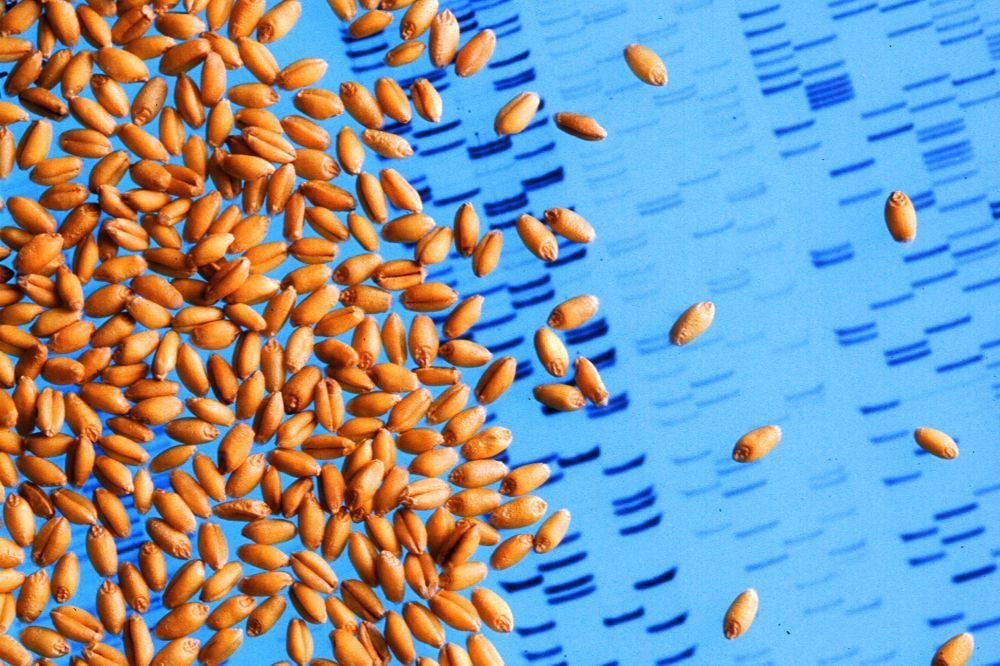

(Jack Dykinga photo courtesy ARS/USDA)
Sequencing the infamously complex genome for bread wheat — a game-changing task for wheat breeding that’s been estimated to take four or five more years — may now just take another couple of years, following a milestone announced Wednesday.
The International Wheat Genome Sequencing Consortium (IWGSC), a team co-led by Canadian researchers, announced Wednesday it has produced a whole genome assembly for Chinese Spring bread wheat.
The assembly “represents a major breakthrough for the IWGSC integrated strategy towards delivering a high-quality reference sequence for each of the 21 bread wheat chromosomes,” project leader Nils Stein, of German plant research centre IPK Gatersleben, said in a release.
Where the IWGSC’s strategy is to study wheat one chromosome at a time, the assembly announced Wednesday was charted using a combination of software, computer programming and bioinformatics tools to look at virtually the entire genome.
The new data can now be integrated with physical-map-based sequence data to produce a “high-quality, ordered sequence” for each wheat chromosome, precisely locating genes and other markers along the chromosomes and providing “invaluable” tools for wheat breeders, the consortium said.
The consortium now expects to have a complete picture of the wheat genome, 17 billion base pairs in all, with a clear idea of how the genes are ordered, within two years.
Given that the wheat genome is five times the size of the human genome, earlier estimates had suggested such work would take four or five more years, the University of Saskatchewan said in a separate release.
The new sequence “is an important contribution to understanding the genetic blueprint of one of the world’s most important crops,” another project leader, Curtis Pozniak of the U of S Crop Development Centre, said in the same release.
“It will provide wheat researchers with an exciting new resource to identify the most influential genes important to wheat adaptation, stress response, pest resistance and improved yield.”
The digital work, he said, has “generated a version of the wheat genome sequence that is better ordered than anything we have seen to date. We are starting to get a better idea of the complex puzzle that is the wheat genome.”
The result, he said, will be greater precision in wheat breeding. “Imagine that you have a blueprint for the order of important pieces of the wheat genome puzzle. With that information, it becomes far easier to assemble the puzzle more quickly into new and improved varieties.”
However, he said, “there is still much work to do to define the function of each of the genetic pieces so that breeders can identify the very best genes in the gene pool.”
While the work was done on just one variety, it’s expected to serve as the backbone to unlock the genetic blueprint for traits in other varieties as well, he said.
The public-private collaborative project, co-ordinated by the IWGSC, is co-led by Stein, Pozniak, Saskatoon researcher Andrew Sharpe of the Global Institute for Food Security and Jesse Poland of Kansas State University.
Project participants also include researchers from San Diego-based genetic sequencing firm Illumina, genomic data firm NRGene, Tel Aviv University and the French National Institute for Agricultural Research (INRA). — AGCanada.com Network